SP 2
Genetic markers predicting side effects, therapeutic response and prognosis in myeloma
One of the main challenges not addressed by the current risk prediction tools are treatment related side-effects, such as neuropathy, bone marrow suppression, or deep-vein-thrombosis. WP2 will assess, for the first time, the association of GWAS data in terms of side effects, such as neurotoxicity, clinical staging, blood chemistry and hematology parameters, bone disease, therapy response, progression free and overall survival. The status of Heidelberg MM genetics as of January 2013 is that 1500 patients with detailed clinical information have been analyzed by GWAS (together with 2300 healthy controls) and among these 1200 are with FISH and 800 with GEP data. This data set will guarantee powerful association analysis between SNPs and each clinical parameter. Ongoing work involves analysis of GWAS data in terms of toxicity emerging in the course of treatment, including neurotoxicity, bone marrow suppression, and thrombosis. Severe side-effects are found in 5 to 20% of patients. Previous studies have suggested genetic basis for neurotoxicity but the present study will be the first GWAS. The GWAS data will also be analysed in terms of the clinical response achieved by treatment, classified as complete, partial or minimal remission and progressive disease.
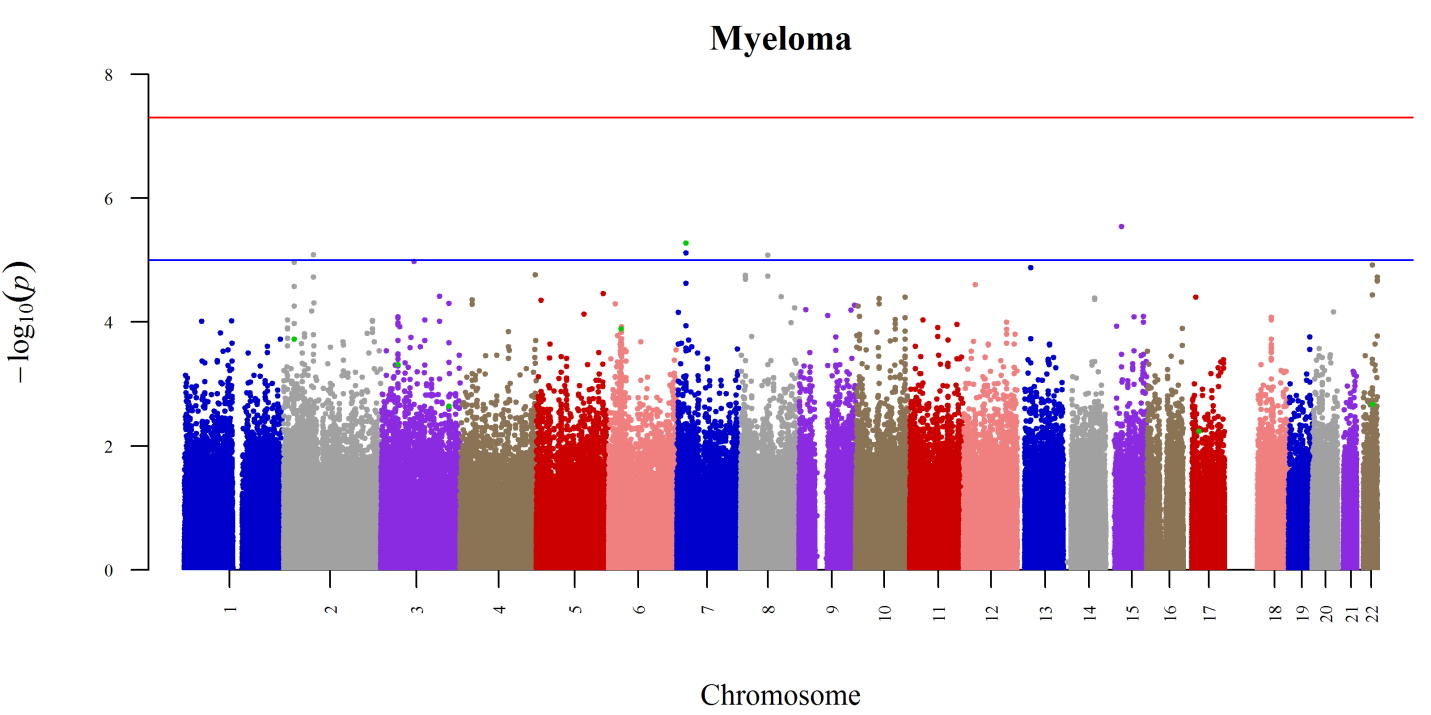
In WP2 further case-control analyses were carried out. In collaboration with WP3 4 additional risk loci for MM were described (Chubb et al., 2013, Nature Genet. 45 (10): 1221-5). These and the 3 previously detected loci were assessed in the precursor disease for MM, MGUS, and in the related disease amyloidosis. They were found to be associated with risk (Weinhold et al., 2014, Blood. 123 (16): 2513-7; Weinhold et al Leukemia in press).



