SP 5
PWAS for identification of functional SNPs and key molecules in arterial injury
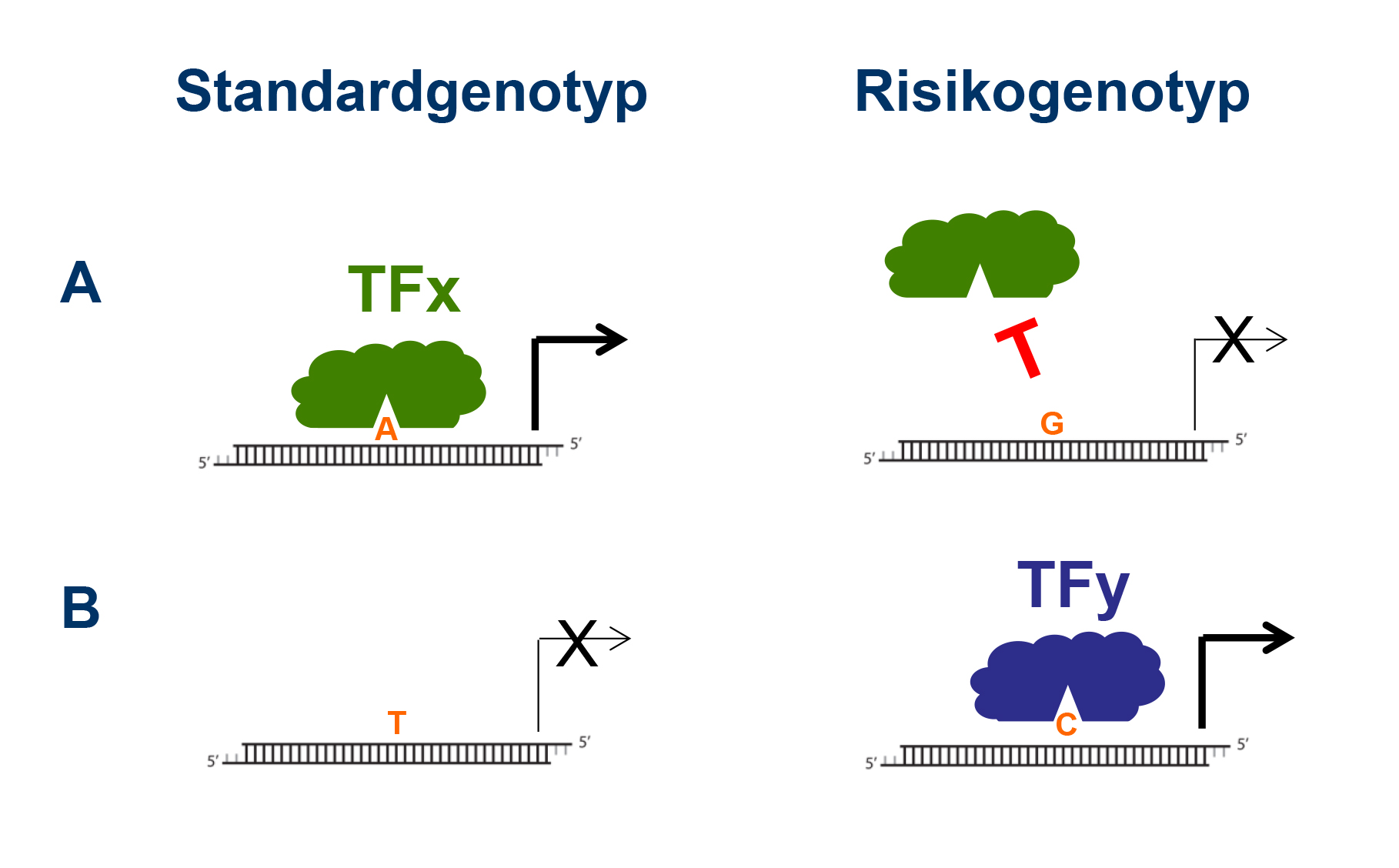
Most hereditary cardiovascular risk factors are genetic variants that are derived from the exchange of a single nucleotide in the genome (single nucleotide polymorphism = SNP), which are found in a fraction of the human population. Most SNPs are located within regions that contain gene regulatory units (rather than in protein-coding regions). Those units are recognized by specific transcription factors that bind to the underlying nucleotide sequence and modify gene expression by interacting with the transcriptional machinery. As a consequence SNPs can either disrupt binding of transcription factors or lead to new binding sites. In both cases gene expression is either increased or decreased, which finally can influence cell function.
In order to unravel SNP-mediated differential transcription factor binding, in this subproject we are conducting a proteome-wide analysis of atherosclerosis-associated SNPs (PWAS). Furthermore, we are planning to study selected SNPs in more detail to understand their involvement in atherogenesis. Our main focus lies on two SNPs, which are located in the regulatory regions of the genes HDAC9 and ADAMTS-7. HDAC9 is an enzyme in the cell nucleus, which itself is involved in the regulation of genes. ADAMTS-7 is a protein, which is located in the extracellular matrix, a structure that is composed of proteins, sugars and signaling molecules. In genome-wide association studies both SNPs were previously found to be significantly linked to coronary artery disease. Meanwhile, we could show that switching off HDAC9 in the mouse leads to an attenuation of atherosclerosis. Similarly, first investigations with mice that do not express ADAMTS-7 showed reduced vessel remodeling after vessel injury. Now our focus is to understand, how both genes are regulated by their underlying SNPs and which molecular function both proteins have during atherogenesis.
The final goal of this subproject is to decipher new molecular signaling pathways in order to find appropriate targets for diagnosis and pharmaceutical treatment of atherosclerosis.
Keywords: atherosclerosis, atherosclerosis single nucleotide polymorphism, SNP



