SP 3
Multi-level analysis of gastric cancer data
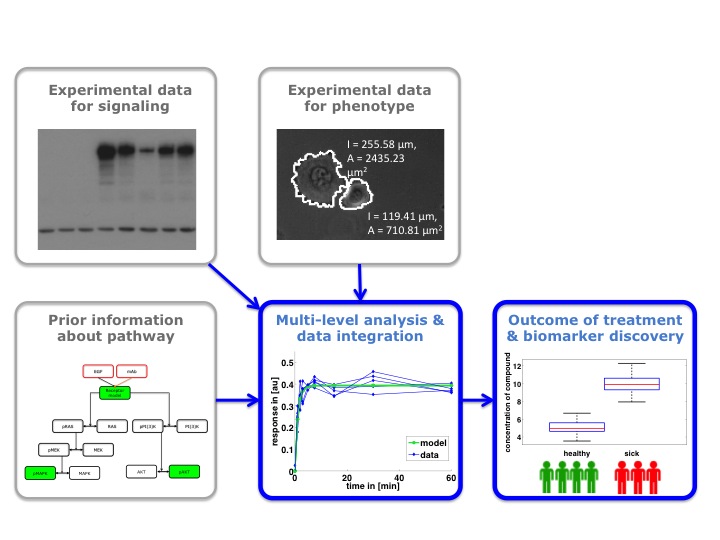
The consortium’s key goal is to understand the connection between molecular mechanisms and the treatment success of HER2- and EGFR-based therapies for gastric cancer. For this, we want to identify and later validate biomarkers for the stratification of patients. The data for this study will be collected experimentally by Subproject 1, 4, and 5 and extracted from literature data by Subproject 2. In this subproject (Subproject 3) we will analyse all available data using classical statistical methods as well as a novel two-step approach developed within the project. This novel two-step approach correlates in Step 1 molecular quantities with the cellular phenotypes, in particular motility, and in Step 2 the cellular phenotypes with the outcome of the treatment. With this combination of high-quality data and tailored statistical methods, we will significantly improve the understanding of gastric cancer.
To achieved the goal of Subproject 3, we will consider the following three work packages:
1) Analysis of correlations between molecular properties and the phenotype based upon available data.
2) Reconstruction of the signalling pathways of cetuximab and trastuzumab in gastric cancer based upon correlation information and available semantic information, and development of a probabilistic model to be integrated in the mobility model.
3) Validation of the reconstructed signalling pathways and phenotype link based upon MALDI imaging mass spectrometry patient samples. Multivariate statistic based extraction and subsequent validation of marker profiles from combined measurement and phenotype simulation data.
Keywords: Stomach cancer, cell migration, metastasis, cell interaction, motility coefficient



