SP 5
Functional characterization of secreted proteins mediating glioma cell invasion
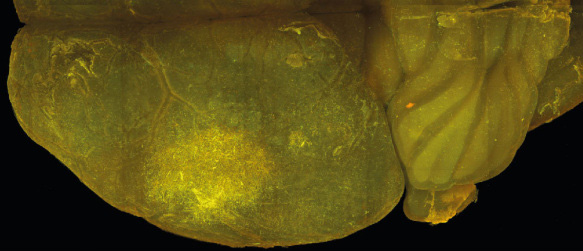
Invasive growth and early infiltration of the surrounding healthy brain is a hallmark of glioma. Glioma invasion was found to be associated with distinct anatomic structures such as the basement membranes of blood vessels as well as the subependyma. Interestingly, glioma cells are also able to use myelin as main trajectory roots, a substrate, which normally
restricts cell migration and spreading. Proliferation and migration of glioma cells are mutually distinct phenotypes: Proliferating cells from the tumor-core do not migrate and, vice versa, migrating cells from the invasion front lack proliferation. This “go-or-grow” dichotomy is a therapeutic challenge since novel treatment approaches mainly target dividing cells.
This invasive nature of glioma cells mainly accounts for their resistance to current treatment modalities. Diffusely infiltrating tumor cells, which evade surgical resection and survive treatment, inevitably give rise to reoccurring tumors. Secreted pro-invasive Extracellular Matrix (ECM)-modulating molecules strongly enhance glioma invasion. These secreted molecules are regulated via the Regulated Ire1-Dependent Decay (RIDD) pathway: Loss of IRE1 signaling results in the upregulation of genes that encode ECM proteins for which the mRNAs were direct targets of RIDD. To date, receptors transducing ECM information and respective downstream signaling molecules driving the RIDD pathway in brain tumors have remained elusive, despite being attractive pharmacological targets.
This subproject aims at identifying substrate-specific ER stress-mediated alterations of secreted proteins mediating brain tumor invasion. It will functionally characterize ER-stress pathways and subsequent changes in secreted proteins for glioma invasion in vitro and in vivo.
Keywords: Unfolded Protein Response, Translational Reprogramming, Translational Regulation



