SP 6
Clinical application of TMM analysis scheme
The aim of SP6 is to apply a standardized telomere maintenance mechanism (TMM) analysis to patient clinical samples, based on the most informative methods derived from the results of subprojects SP3-SP5. The precise methodology (sequencing, automated imaging, qRT-PCR, etc.) will depend on the number and type of parameters that need to be assessed in order to provide the most relevant information. The development of the TMM analysis scheme will be a highly interactive and iterative process with SP3-SP5, involving continuous input of SP6 into experimental assessment and validation through several development cycles. These data, and complementary information produced in the context of other ongoing sequencing studies, will be used to generate a more rational targeted strategy for clinical applications. We will focus on the correlation of the TMM classification with prognostic impact or prediction of therapy response. Since genetically distinct subgroups are associated with very specific TMM patterns, we anticipate that TMM patterns will be an excellent surrogate marker for biological subgroups and thus clinical parameters such as response to therapy.
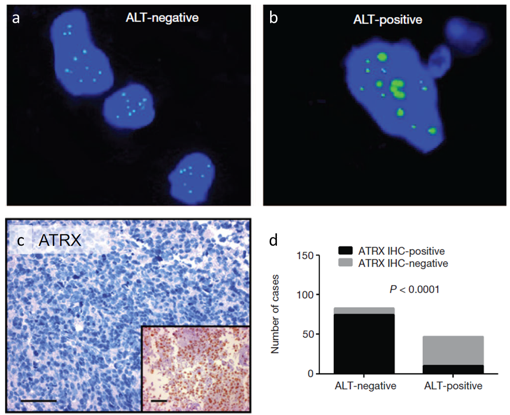
It may be assumed that DNA-damaging therapies like radiation or treatment with alkylating agents might promote the selection of ALT (Alternative Lengthening of Telomeres)-positive cells through increased recombination of telomeres, highlighting potential interactions between TMM and treatment response. The TMM of different tumor samples will be classified for a subsequent correlation with clinical data, in order to make better predictions about prognosis and for the choice of therapy based on tumor-biological markers.
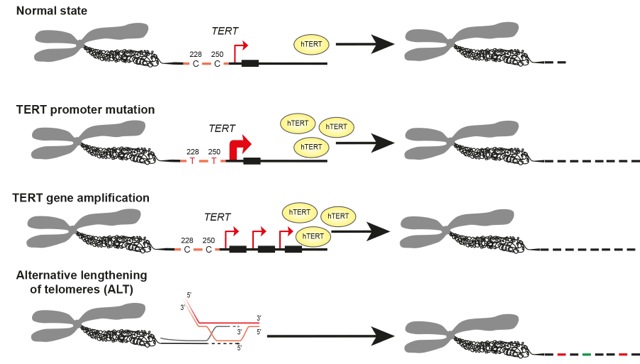
Keywords: Pediatric; cancer; brain; glioblastoma; medulloblastoma; histone; telomere



