SP 8
Characterisation of antibody-antigen interactions and adapation on a diagnostic platform
During the last decades, acute rejection and allograft survival rates have been significantly improved by the introduction of new potent immunosuppressants. The main aim to treat patients with immunosuppressants is to suppress the rejection of the allograft by the cellular and humoral immune systems. Especially understanding and treatment of humoral (antibody-mediated) rejection remains a major challenge. Indeed, emergence of donor specific HLA antibodies (DSA) may hamper the development of operational tolerance in transplant patients. Clinical relevance of DSA may be determined by antibody quantity, complement binding capacities but also binding strength to particular epitopes.
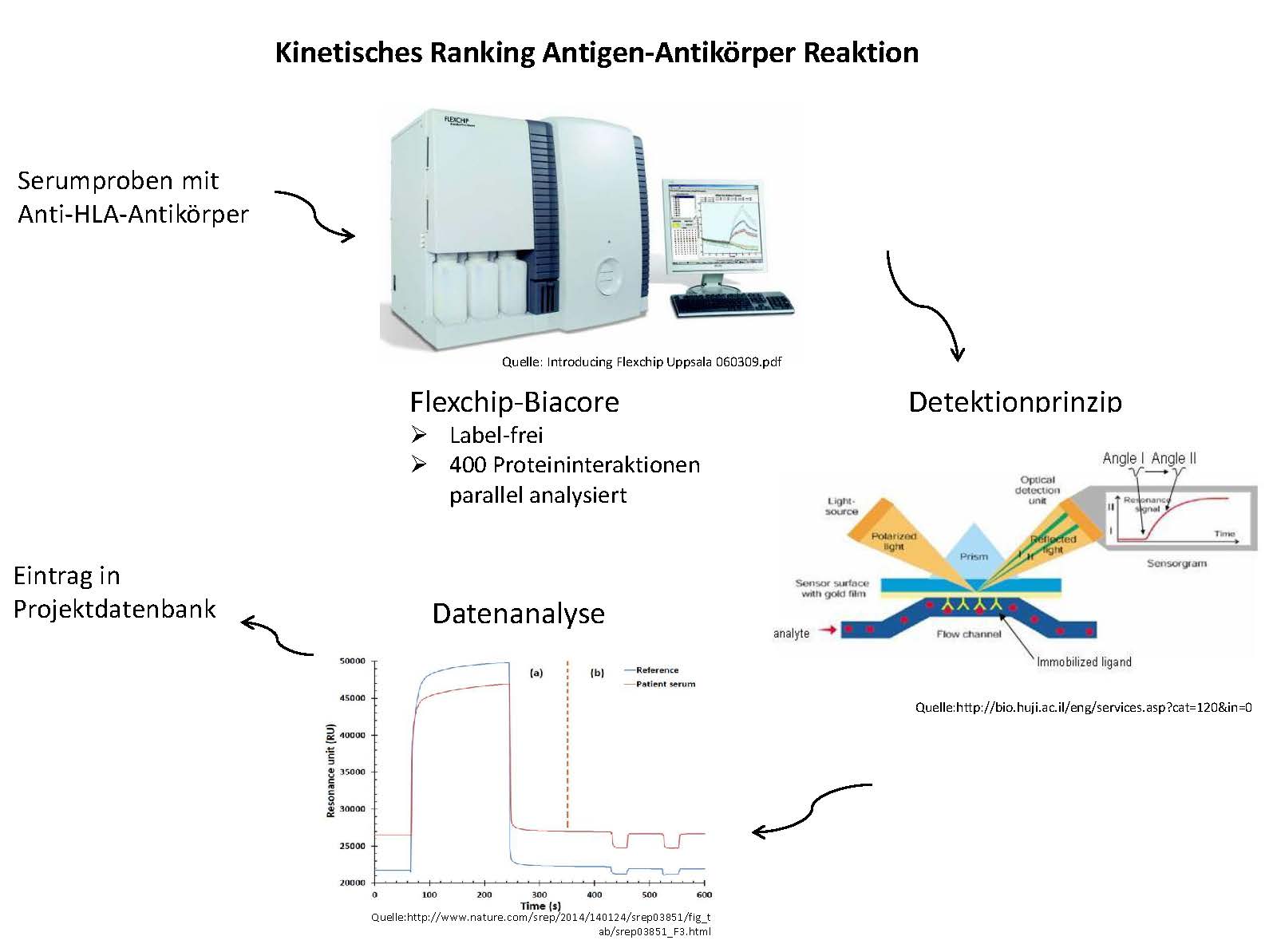
Previously we could show by an in vitro assay that different IgG and IgM antibodies have different affinities although they recognize identical targets.
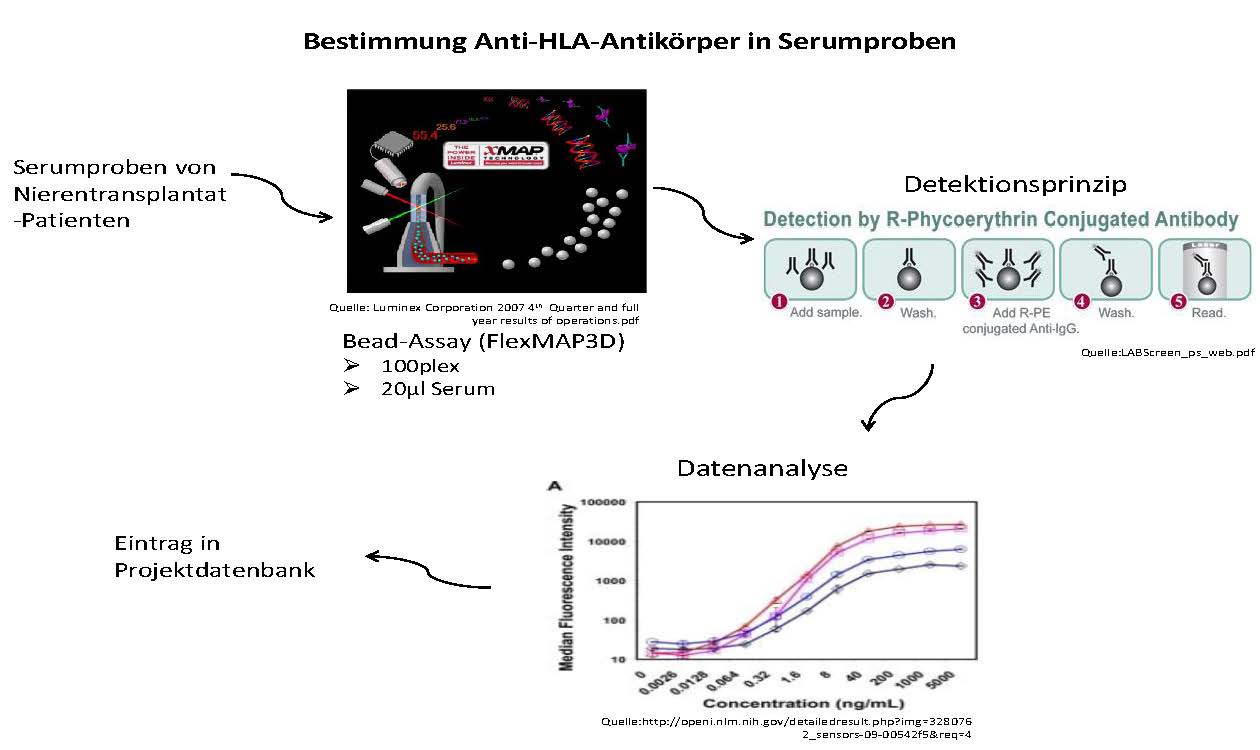
We will use the serum samples of the well-characterized patient cohort of 500 patients provided by the partners to analyze and characterize contained anti-HLA antibodies binding in vitro. Besides the anti-HLA antibody and particular DSA concentration, the binding affinity and epitope specificity will be determined. All generated data will be provided to bioinformatics partners for data integration and mathematical modelling. The proposed system medicine based established model will be used to perform a prospective risk-adjusted clinical trial on personalized treatment in kidney transplant patients.